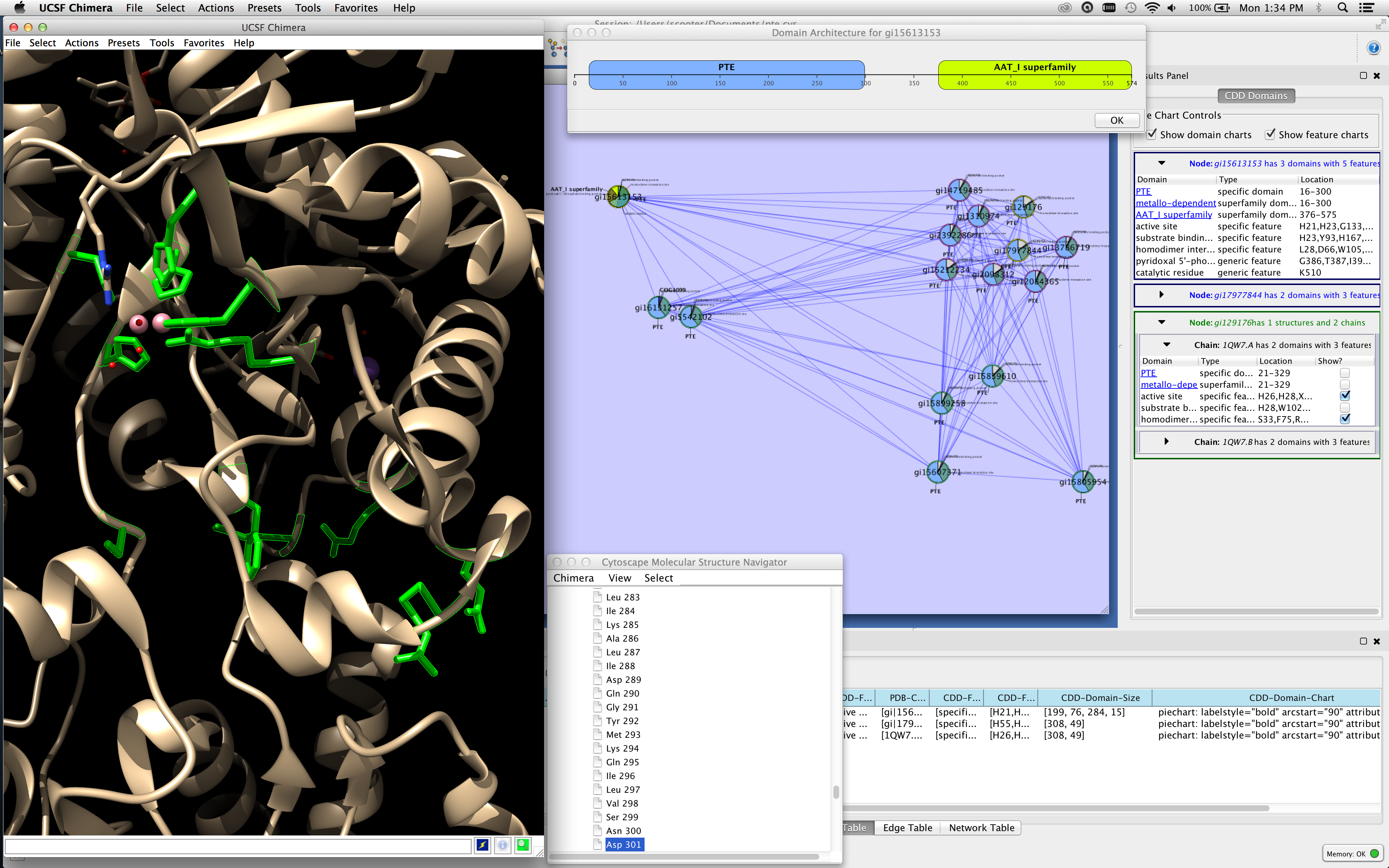
Since it was created in July 2013, CytoCluster has been downloaded more than 9700 times in the Cytoscape App store and has already been applied to the analysis of different biological networks. Module (cluster) detection in PPI network. This clustering method was achieved by the build-in interface ‘scanpy.tl.leiden’ from ScanPy, with the parameter ‘resolution’ equal to 0.8.
CLUSTERING APP CYTOSCAPE SOFTWARE
CytoCluster can be easily expanded, so that more clustering algorithms and functions can be added to this plugin. Search for posts about graph clustering Ask a question about graph clustering. Cytoscape: a software environment for integrated models of biomolecular interaction networks. In addition, BinGO is used to determine which Gene Ontology (GO) categories are statistically overrepresented in a set of genes or a subgraph of a biological network. Type 1 diabetes mellitus (T1DM) is a metabolic disorder for which the underlying molecular mechanisms remain largely unclear. Moreover, visualization of different topological variation of nodes and clusters over time enables users to quickly identify the most variations across many. The main function of these six clustering algorithms is to detect protein complexes or functional modules. Users can select different clustering algorithms according to their requirements. Here we present CytoCluster, a cytoscape plugin integrating six clustering algorithms, HC-PIN (Hierarchical Clustering algorithm in Protein Interaction Networks), OH-PIN (identifying Overlapping and Hierarchical modules in Protein Interaction Networks), IPCA (Identifying Protein Complex Algorithm), ClusterONE (Clustering with Overlapping Neighborhood Expansion), DCU (Detecting Complexes based on Uncertain graph model), IPC-MCE (Identifying Protein Complexes based on Maximal Complex Extension), and BinGO (the Biological networks Gene Ontology) function. Furthermore, the visualization of clustering results is crucial to display the structure of biological networks. The constructed MSC tree is consistent with the best phylogenetic models from the character-based approach.Nowadays, cluster analysis of biological networks has become one of the most important approaches to identifying functional modules as well as predicting protein complexes and network biomarkers. An example is the clustering of a 46-coronavirus network in three characteristic levels as shown in the screenshots. Here, we have demonstrated that the constructed MSC tree provides phylogenetic information for complex systems from a distance-based approach. We have developed the MSClustering app in Cytoscape for an automated, efficient, and hierarchical clustering of complex networks without any input parameter. As many important real-world clustering/classification problems are intrinsically hierarchical, it is desired to develop an efficient clustering tool for the automated clustering of complex systems at various characteristic levels. It is now integrated with Cytoscape and provides immediate visualization and statistical analyses.Ĭlustering/classification is an important step in understanding the present diversity and past evolutionary history of a complex system. mir-221-3p/mir-222-3p have the same seed sequence and were considered together as a cluster. Therefore it is efficient (computation time is linear in N for N < 1500) and has been successfully applied to study various systems in biology, scientometrics, literature analyses, and finance. Molecular interaction network to common X-linked ID genes and miRNAs by Cytoscape 3.6.1.
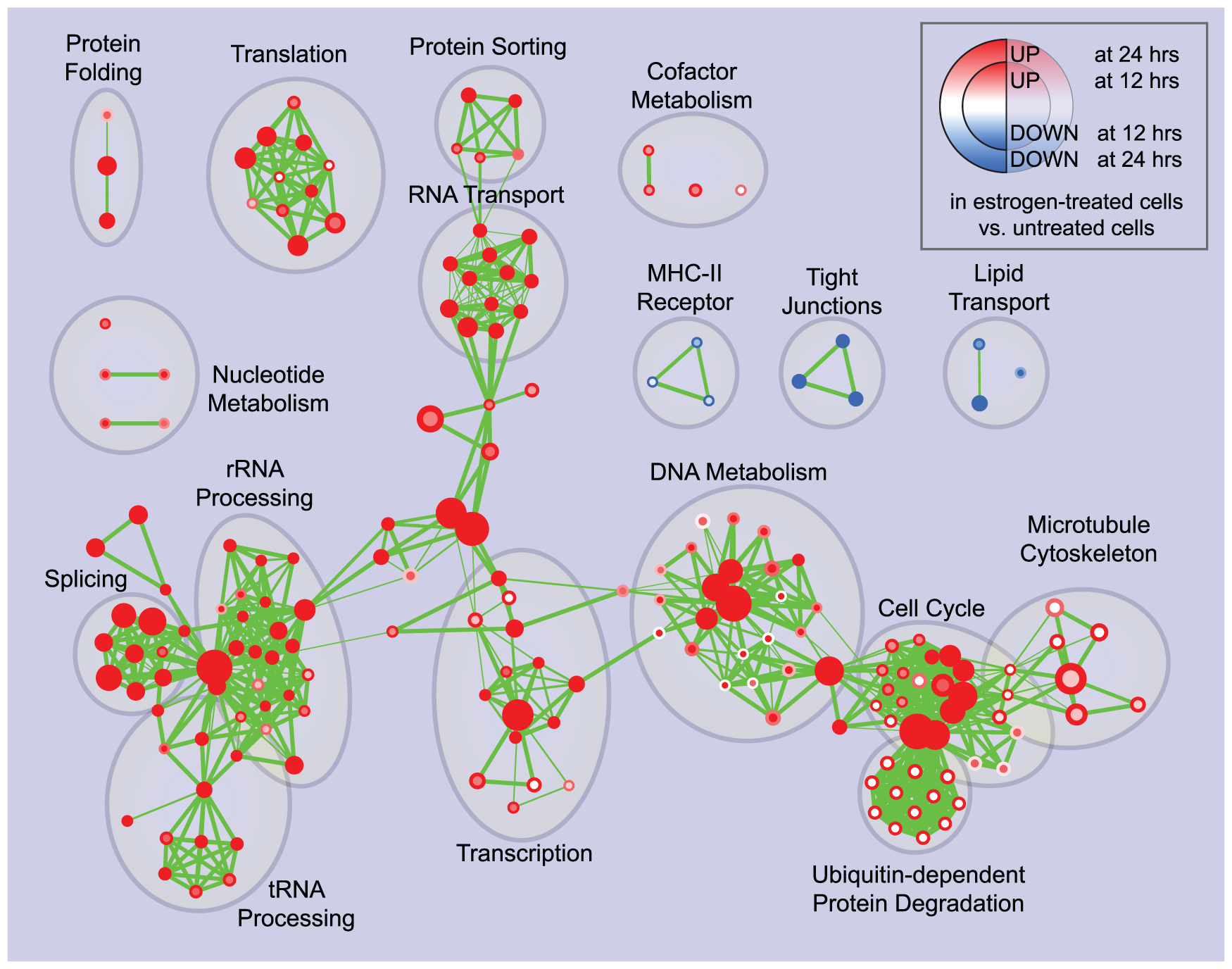

For a system of 2500 nodes, it takes about 36 seconds to perform a hierarchical clustering of the system. It has a system size dependence in the memory of storage and operations of the algorithm. After transforming the N^2-distance matrix into an N-list of shortest connections for a system of N nodes, MSClustering groups the system hierarchically for several characteristic levels of resolution, without the input of parameters.
CLUSTERING APP CYTOSCAPE INSTALL
MSClustering is a Cytoscape tool for multi-level clustering of complex networks and provides immediate visualization and analyses in the Cytoscape platform. Start Cytoscape, install the CytoCluster.jar through App Manager (Apps App Manager Install from file.) 2.
